DISSECTING ENHANCERS AT NUCLEOTIDE RESOLUTION
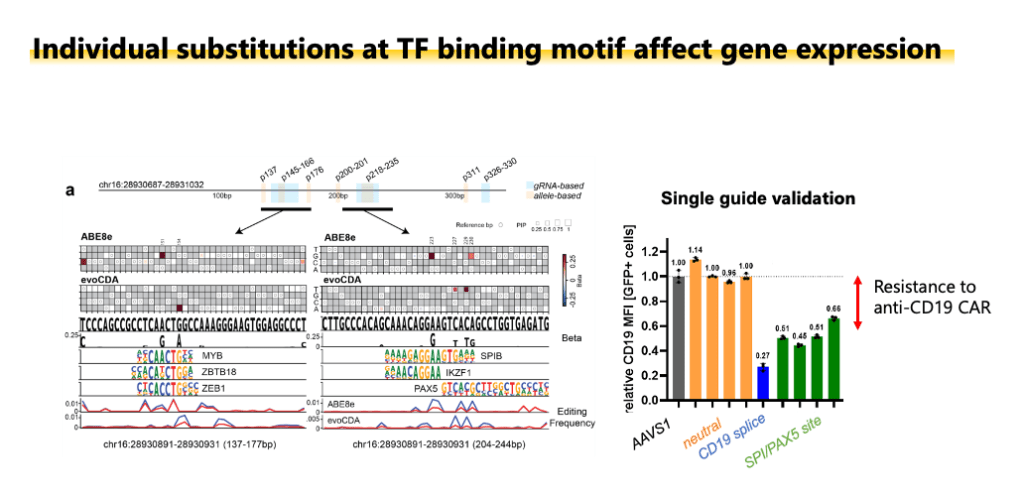
Enhancer scanning by dense base editing coupled with allele sequencing. Adapted from Becerra, Wittibschlager, Patel et al. 2026 Nature Communications.
Functional non-coding regions such as enhancers and insulators play an essential role in the regulation of cell type-specific gene expression programs, and mutations at non-coding elements can drive observable phenotypic changes comparable to those driven by mutations at coding sequences (see Seruggia et al. 2015; Seruggia et al. 2020). However, due to lack of appropriate technology, the grammar of enhancers, and the mechanism of disease-associated variants remain unsolved. In our lab, we focus on germline variants predisposing to pediatric leukemia, that often occur at putative enhancers. What is the mechanism of sequence variants at enhancers and how are these associated with leukemia? We use epi/genome editing (CRISPR, CRISPRi, CRISPRa, base and prime editing), chromatin profiling (ChIP-seq, Cut and Run, ATAC-seq) and computational biology to answer these questions. See Becerra, Wittibschlager, Patel et al. 2026 on dense base editing at enhancers.
MECHANISM AND VULNERABILITIES OF THE SAGA COMPLEX IN LEUKEMIA
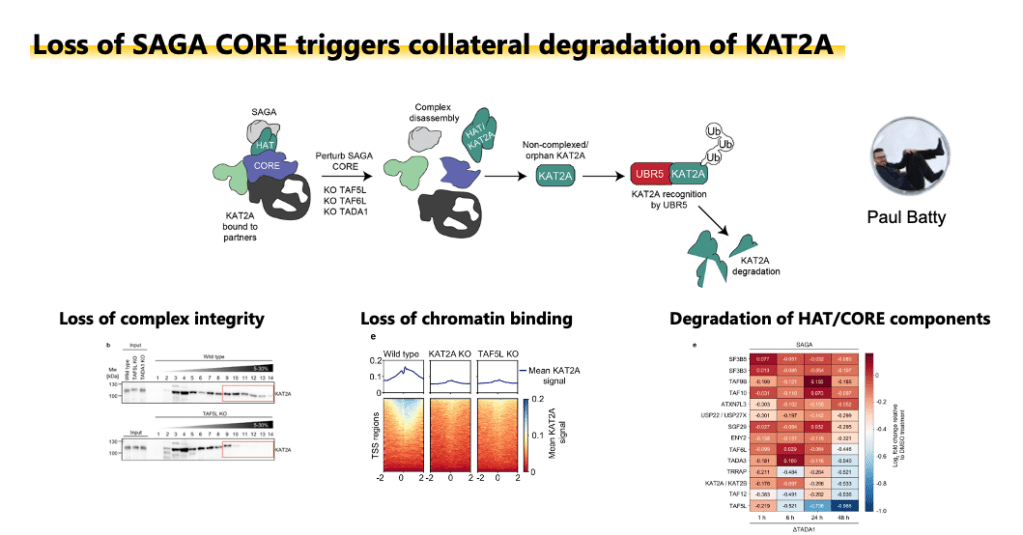
Targeting components of the CORE module of SAGA leads to complex disassembly and KAT2A degradation. Adapted from Batty et al. 2026 Nature Communications.
The SAGA complex regulates gene expression by deposing H3K9ac at promoters. We previously reported that loss of structural components lead to global loss of histone acetylation in mouse embryonic stem cells (Seruggia et al. 2019). In our recent paper (Batty et al. 2026) we showed that the integrity of the SAGA CORE module maintains the stability of the histone acetyltransferase KAT2A. Disruption of CORE components converts KAT2A into an orphan substrate, triggering degradation by the ubiquitin-proteasome and reducing histone acetylation, revealing a proteostatic vulnerability in SAGA-dependent systems. Motivated by these findings, we are currently investigating how to target components of the SAGA complex in the context of leukemia.
MOUSE MODELS

We are interested in developing new animal models to study hematopoiesis and leukemia development. In the lab, we design and prepare CRISPR reagents suitable to treat mouse embryos at the 1-cell stage. We have successfully targeted regulatory sequences (see Seruggia et al. 2015, Seruggia et al. 2020, Singh et al. 2022), introduced specific point mutations (see Sher et al. 2019) and generated degrons or conditional alleles for our favorite genes (see Mehta et al. 2022; Manieri et al. 2023).
Values of the lab: we enjoy working on curiosity- and technology driven projects. We thrive to identify new molecular mechanisms through deep dives in biology. Our mission is to understand disease better and identify new actionable cancer vulnerabilities.
